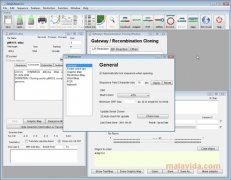
Support for Additional File FormatsĮnhancements to Outlining Shared Domains in Aligned Sequences Searching for CRISPR PAM SitesĪ new tool Analyze | CRISPR PAM sites… menu item will scan Nucleic Acid sequences for Protospacer Adjacent Motifs associated with the CRISPR Cas9 and related enzymes cleavage and modification functions. Plus MacVector 18.1 will be released very soon with full Apple Silicon Universal Binary support (ARM64 and X86_64). MacVector 18.0 is fully supported on El Capitan to macOS Big Sur. Finally, there are major enhancements to the protein multiple sequence alignment visualization of domains and the usual macOS compatibility enhancements. There is a new CRISPR PAM sequence searching function and a Trim by Quality option for Sanger sequence files. MacVector 18 adds a number of new functions, including the ability to read several requested file formats such as Sequencher assembly project files along with Serial Cloner and SnapGene sequence files. MacVector 18.1 is fully supported on macOS Sierra to macOS Big Sur. The use of a comma as the decimal point separator is now supported throughout MacVector for international localization. In addition, small deletions are now reported, again using standard nomenclature. The SNPs tab has been revised to use standard reporting for both nucleic acid SNPs and for corresponding changes to amino acid sequences in CDS features. The align to reference algorithm has been slightly tweaked to do a better job of aligning reads that have insertions relative to the reference. Improved Align to Reference SNP Reporting The main application icon and window toolbar appearance have been updated to match the macOS Big Sur “look and feel”. Other than the ability to run natively on M1 processors, MacVector 18.1.1 is essentially identical to MacVector 18.0.1 with the exception that the embedded Python framework has been updated to version 3.9. MacVector 18.1 is a Universal Binary, meaning that it can run natively on both Intel and Apple Silicon Macintosh computers. MacVector 18.1 is a Universal Binary application that means it runs natively on Apple Silicon and Intel Macs. To reduce clutter in the Assembly Project window toolbar, all of the assembly algorithms have been consolidated into a single Assemble toolbar button with a dropdown menu. This functionality replaces the old Primer Converter utility. You can import primer data into a MacVector Primer Database (.nsub) file from an Excel or CSV formatted file. Importing of Primer Databases in TSV or CSV Format Where a window has a context-sensitive menus available these views now contain a “hamburger” button (three parallel horizontal lines) that displays the same context sensitive menu when clicked on. There is a new Remove Gaps context-sensitive (right-click) menu option that deletes residues in reads that correspond to a gap in the consensus sequence. When aligning ABI chromatogram data, or plain sequences, the Map tab now graphically displays the “trimmed” regions at either end of the sequences. In addition, the Sensitivity setting can now be lower due to the enhanced consecutive gap detection, which also speeds up calculations. The alignment algorithm has been further optimized for speed. The Align to Reference alignment algorithm has been overhauled to do a much better job handling larger numbers of gaps in the alignment between a reference sequence and a read. It is supported on macOS Sierra (10.12) to macOS Monterey (12). MacVector 18.2 is a Universal Binary that runs natively on both Apple Silicon and Intel Macs. As usual there’s been a lot of bug fixes and changes that you probably do not care about! But be assured that everything we do makes MacVector a future proof and modern macOS application that you can rely on!. 
Importing Sequencher project (.SPF) files has been significantly enhanced.Long sequencing reads from PacBio and ONT sequencers can now be assembled in Align to Reference. You can also basecall heterozygotes in a trace file in the Single Trace Editor. The tool works on multiple trace files in the Assembly project manager or the Align to Reference editor. You can also analyze Sanger trace files and permanently change the basecalled sequence with an IUPAC ambiguity code representing the called heterozygote. The heterozygote analysis tool analyzes one or multiple Sanger trace files and reports on all possible heterozygotes. Heterozygote Analysis of Sanger trace files It is supported on macOS High Sierra to macOS Ventura and is a Universal Binary that will run natively on Apple Silicon Macs as well as existing Intel Macs. MacVector 18.5 was developed and tested on macOS Ventura. ( See here for a detailed history of releases) MacVector 18.5 Overview
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
